Portfolio
Regularized canonical correlation analysis (rCCA)
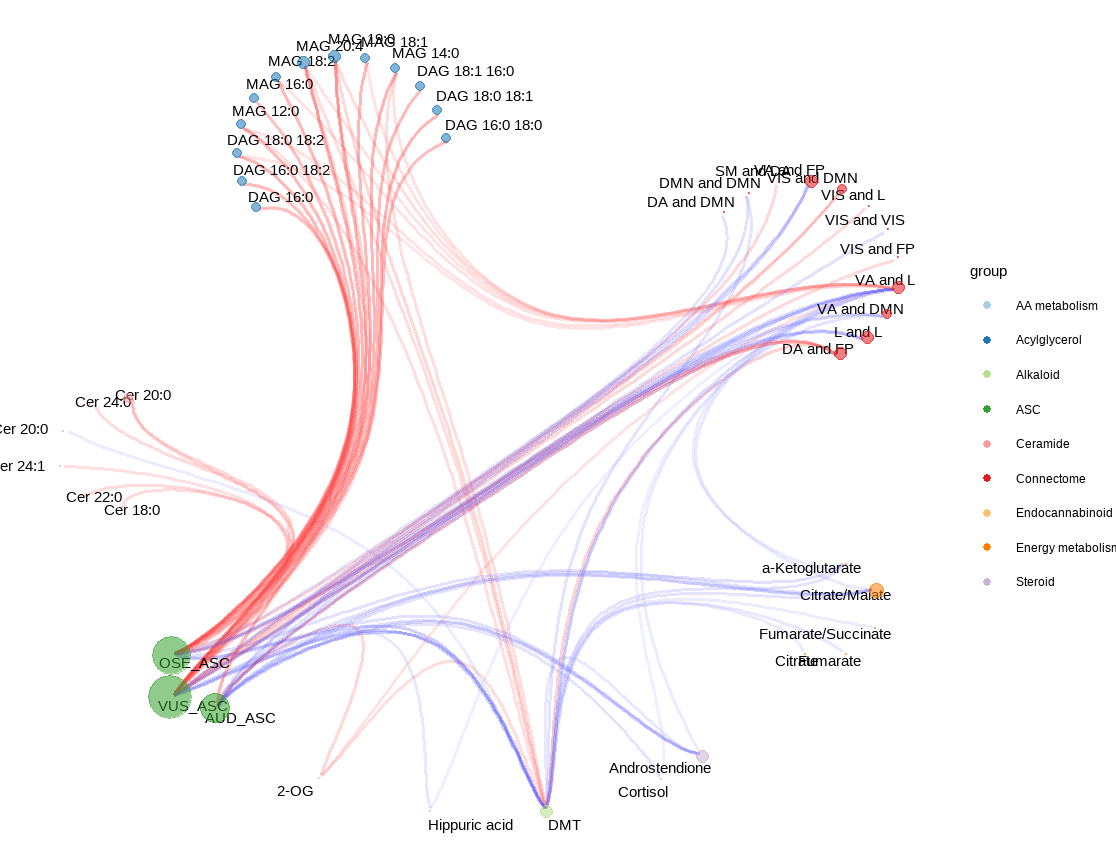
Regularized canonical correlation analyses (rCCA) were performed on all individuals using the regularization of the empirical covariance matrices. This model serves as an integrative multivariate approach to assess correlations between metabolomics, plasma alkaloids, and drug subjective effects. Lambdas were estimated utilizing the Shrinkage method. Next, integrative pathway network and the corresponding clustered heat map were built from the rCCA outputs. Clustered heat map dendrograms were computed from hierarchical cluster analysis, using the Euclidean distance and the complete linkage method.
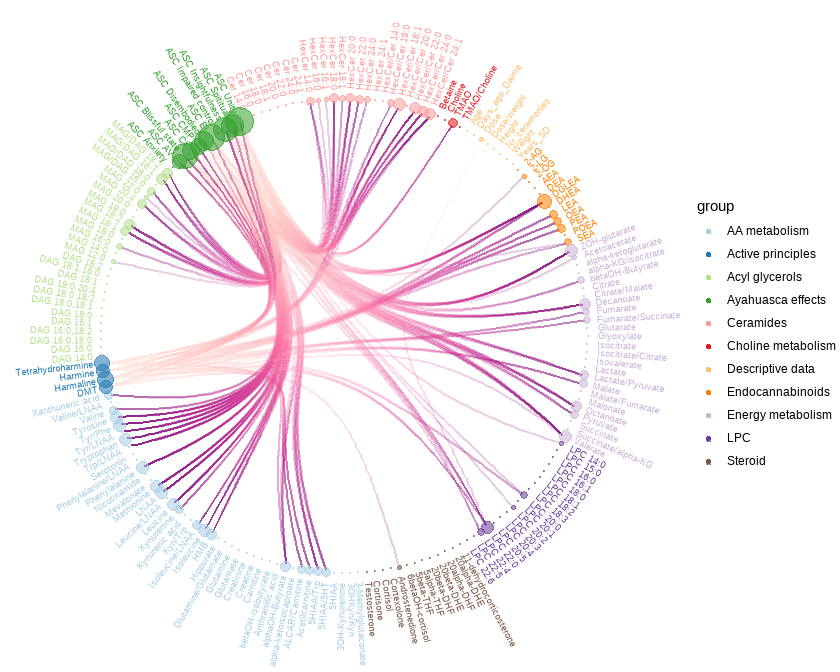
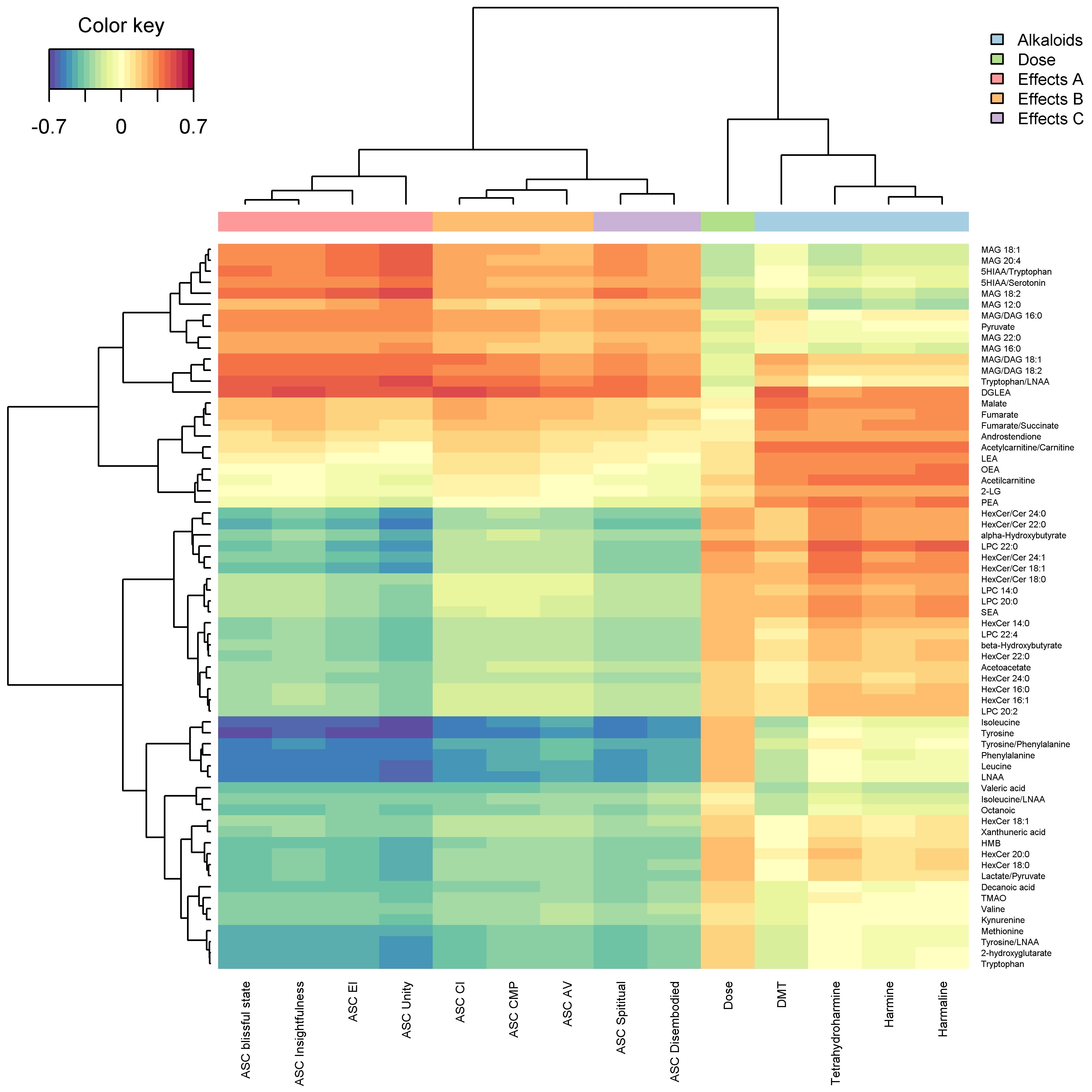
Link to conectome-metabolome network after the administration of a psychedelic drug (link can take a while):
https://metabolomeconnectome.netlify.app/
Previous work:
https://doi.org/10.1016/j.biopha.2022.112845
Multiblock data analysis
N-integrative framework models that allow to integrate multiple datasets while explaining their relationship with an outcome (working on this).
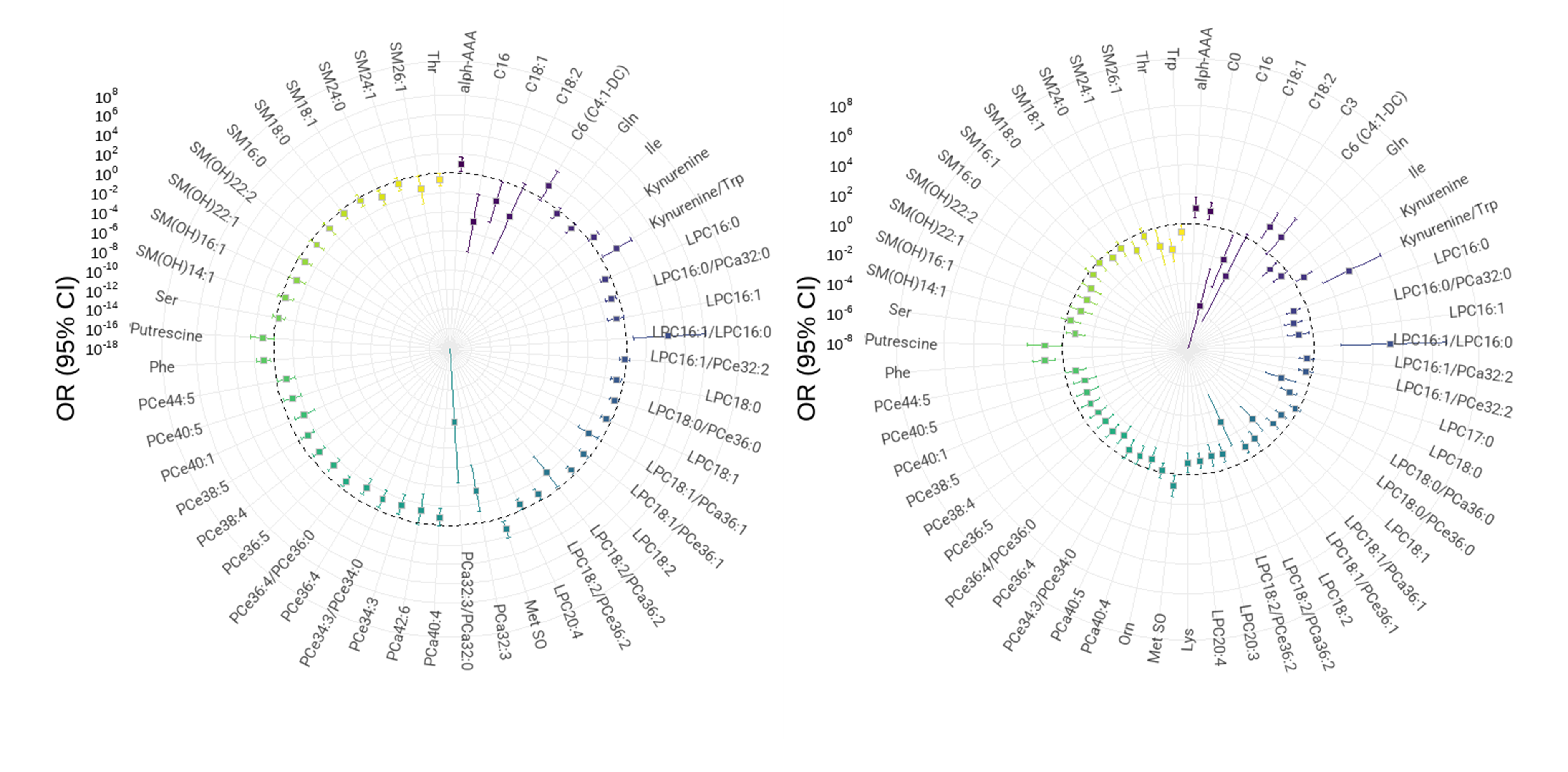
Odds ratios polar plot
Using odds ratios (ORs) and generalized, mixed, logistic or poison models, we can measure how strongly an event is associated with exposure. Probability ratios are comparing two sets of odds: the odds of an exposed group experiencing the event versus the odds of a non-exposed group experiencing the event. In case-control studies, odds ratios are commonly reported. The odds ratio helps identify how likely exposure is to lead to a specific event. The larger the odds ratio, the higher odds that the event will occur with exposure. Odds ratios smaller than one imply the event has fewer odds of happening with the exposure.
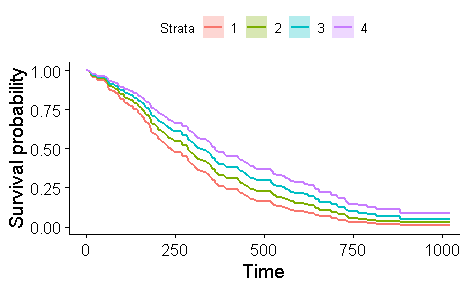
Kaplan-Meier survival analysis
The Kaplan-Meier estimate is the simplest way of computing the survival over time in spite of all these difficulties associated with subjects or situations. For each time interval, survival probability is calculated as the number of subjects surviving divided by the number of patients at risk.
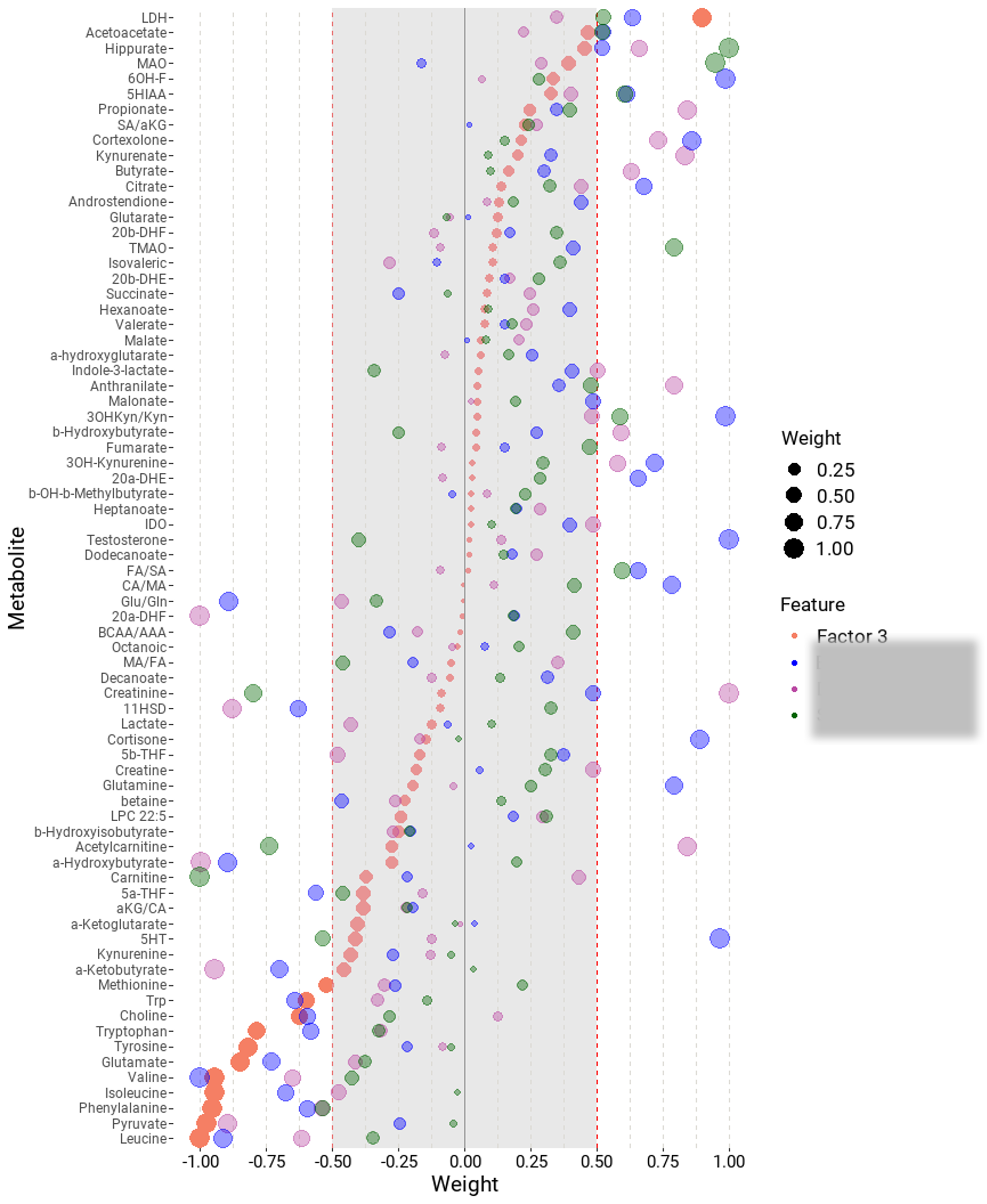
Factor analysis with MOFA
Multi-Omics Factor Analysis
MOFA is a probabilistic factor model that aims to identify principal axes of variation from data sets that can comprise multiple datasets layers and/or groups of samples. The algorithm also allows for multimodal analysis. Additional time or space information on the samples can be incorporated using the MEFISTO framework.
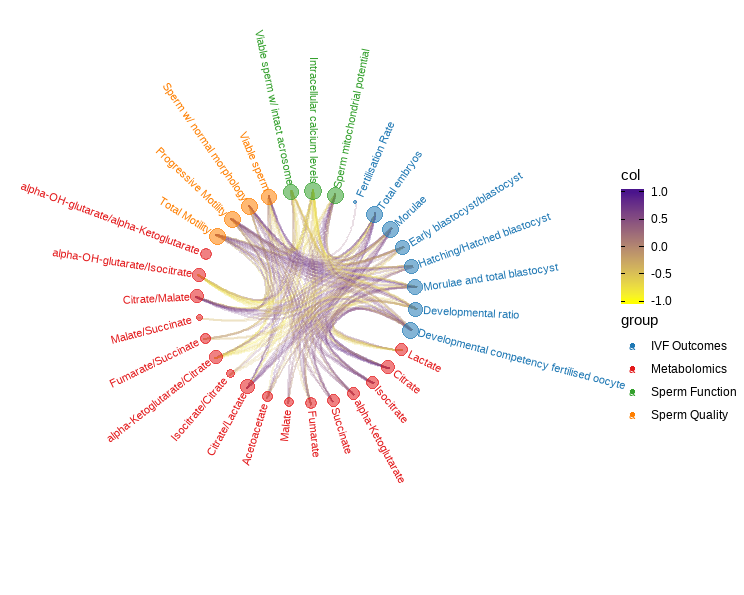
Multi-block PLS: data integration
N-multi-block with Projection to Latent Structures model
Integration of multiple data sets measured on the same observations were carried out utilizing the multiblock integration with Projection to Latent Structures model. This model was performed to assess multiblock correlations between sperm quality, sperm function, IVF outcomes, and metabolomic blocks from the same observational units. Pair-wise similarity matrix was obtained from the two correlated latent components obtained through the projection to latent structures method. A relevance network graph was created to describe connections between the four datasets.
https://www.nature.com/articles/s42003-023-04715-3
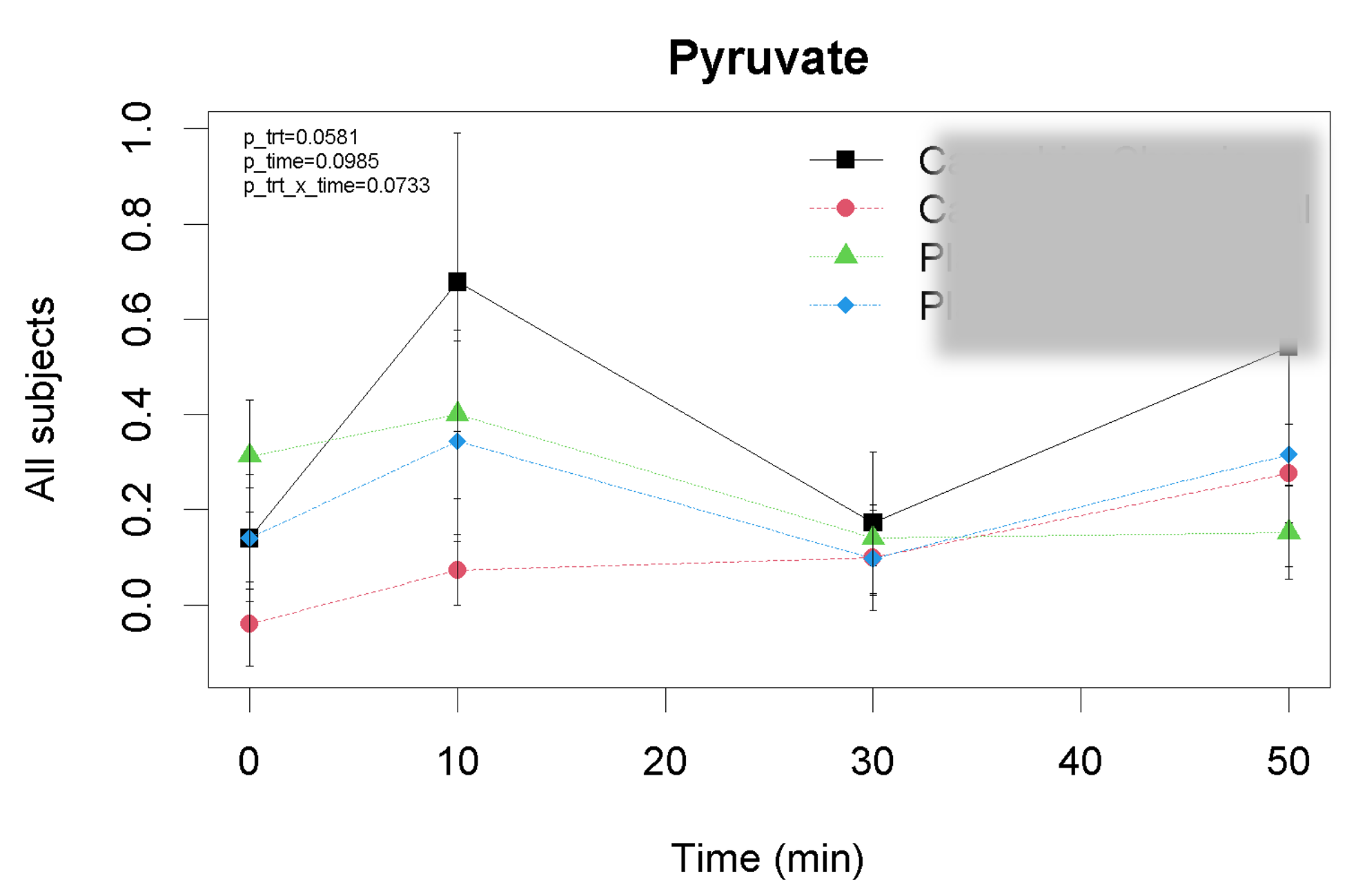
Kinetics
Kinetics modeling of plasma pyruvate after a drug/placebo treatment. By utilizing a generalized mixed linear regression with an autoregresive term, we can assess differences between treatments within the first hour.
(working on this Portfolio page)







